Codon | Where the Genes Flow
Awards & Nominations
Codon has received the following awards and nominations. Way to go!
The Challenge | Where the Genes Flow
GeneX
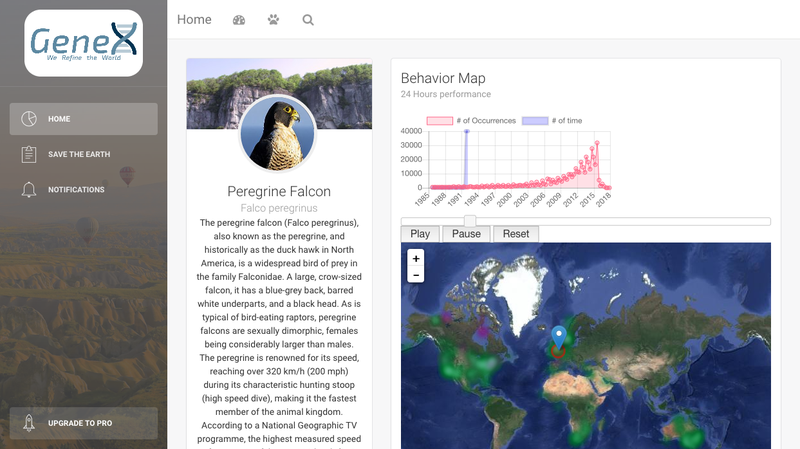
GeneX is a analyzer and visualizer on gene flow and local adaption patterns
Background:
FACTORS (BARRIERS AND FACILITATORS TO THE GENE FLOW)
Currently, our team has identified several general factors which govern the gene flow of any given species.
- Climate changes
- Migration patterns
- Environment and Landscape
- Human behaviours
- man-made artificial barriers
- Deforestation
- War Zones
Research:
ABSTRACT
Two main transfer methods were identified,
- Vertical gene transfer - Transfer of genetic material is from parents to offspring through sexual or asexual reproduction.
- Horizontal gene transfer -Movement of genetic material from a donor organism to a recipient organism that is not its offspring.
According to the preliminary research our team executed, these two transfer methods are the cause of gene flow between different populations of the species. However, our team mainly focused on the vertical flow which is more feasible to track in the given time interval.
Challenges faced:
According to many research, tracking a species in order to experiment on the gene flow is not quite easy and feasible. As a result many of the experiments are involved in accruing a proper estimate for the gene flow. So the main challenge we faced was how to efficiently estimate a gene flow with data in hand.
New frontier:
We team codon presents a new approach to evaluating the gene flow based on the migration patterns of any given species.
According to the research, Nm (the product of the effective population number: N and rate of migration among populations: m) is the factors that act as the direct measure of gene flow. So we thought of using migration pattern data globally available via Nasa portals and other resources such as GIBF biodiversity data. This method might open new research on how to utilize machine learning in order to evaluate and estimate gene flow via a probability value.
Detailed steps of our method are as below:
- Gathered the migration and mobility data of the species and prepared a sample dataset.
Link to data-set: https://github.com/TeamCodon/geneflow_detector/tre... - Use a clustering technique (Machine learning technique) to divide those geospatial data into clusters.
- We exploited the K-Means clustering method with a predefined number of clusters. This can be optimized further.
- Then evaluate the boundaries of those clusters and get the density of the cluster intersections as a measure of the probability of gene flow.
Future we see:
- This method can be used as a direct and more accurate way of measuring gene flow of any given species.
- This method directly shows the correlation between migration pattern and the gene flow.
- Clustering method can be improved further.
- Use gene flow to predict population growth and risks.
GeneX prototype can be found here: https://teamcodon.github.io/genex_webapp/
Github link : https://github.com/TeamCodon
Resources:
gbif.org - Occurrences of species.
Movebank.org - Species tracking data
Technologies:
Front-end:
- Java-script - Leaflet.js, Leaflet.heat
Machine Learning (clustering):
- Python - scikit-learn
Reference
- Ellstrand, Norman C. "Is gene flow the most important evolutionary force in plants?." American journal of botany 101.5 (2014): 737-753.
- Mallet, James. "Gene flow." SYMPOSIUM-ROYAL ENTOMOLOGICAL SOCIETY OF LONDON. Vol. 20. 1999.
- Sexton, Jason P., Sandra B. Hangartner, and Ary A. Hoffmann. "Genetic isolation by environment or distance: which pattern of gene flow is most common?." Evolution 68.1 (2014): 1-15.
- Larson, Allan, David B. Wake, and Kay P. Yanev. "Measuring gene flow among populations having high levels of genetic fragmentation." Genetics 106.2 (1984): 293-308.
SpaceApps is a NASA incubator innovation program.